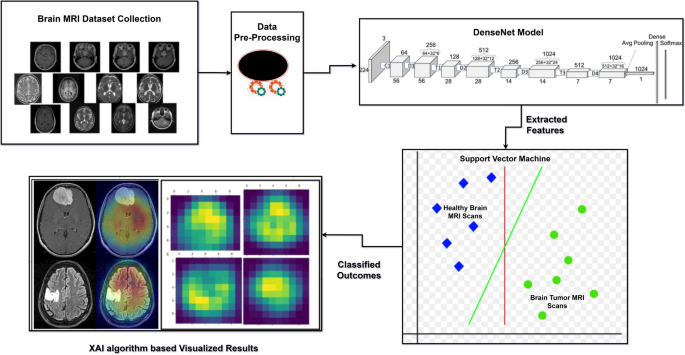
The methodology presented in this study outlines the development of an automated hybrid model that integrates the DenseNet convolutional neural network architecture with a Support Vector Machine (SVM) classifier. To enhance model transparency and clinical interpretability, the architecture is further combined with an Explainable Artificial Intelligence (XAI) algorithm—specifically, Gradient-weighted Class Activation Mapping (Grad-CAM). This comprehensive system is designed for the detection and classification of brain tumors using MRI imaging data. The objective is to support medical professionals in making more accurate and timely diagnoses while overcoming several challenges posed by conventional techniques, such as limited generalization, lack of interpretability, and suboptimal classification accuracy. The proposed hybrid approach demonstrates improvements in diagnostic precision, trustworthiness, and outcome reliability. The complete workflow of the methodology is illustrated in Fig. 1.

Methodology for detection of brain tumor.
Problem definition and data acquisition
The first part of the research process is to properly define the problem, in this case the use of artificial intelligence to the proper classification and early diagnosis of brain tumors. Brain tumors are one of the deadliest neurological diseases, and early diagnosis is crucial to maximize the effectiveness of treatment. To address this diagnostic problem, the proposed study would employ a sophisticated machine learning approach to classify MRI information into two basic classes: healthy and brain tumor. The second process, subsequent to problem specification, is the acquisition of suitable data that can be used for training as well as verifying the model. The data employed in this research comprises brain MRI images as depicted in Fig. 2 from publicly available medical image databases. These databases offer pre-classified as well as annotated MRI scans38 comprising both normal and cancerous brain images. The information includes MRI images in various formats such as JPEG and DICOM, with varying resolutions and pixel intensity values, which need a stringent preprocessing phase before training. The diversity and range of the gathered dataset are intended to enable the model to learn from a wide range of real-world presentations of brain tumors and thus enhance its generalizability to new cases.

Representation of brain MRI scans: (a) Healthy, (b) Brain tumor.
Dataset description
The data used in the current study were taken from the publicly available Kaggle data repository, i.e., the dataset created by Sartaj38. This dataset contains a total of 3264 brain MRI images in two classes: ’Tumor’ and ’No Tumor’. The images are grayscale MRI slices provided in .jpeg format with different resolution, brightness, and orientation. The class distribution has 1683 images marked as tumor and 1581 as no tumor, which is a relatively balanced set appropriate for binary classification problems.
Each image preserves the structural brain anatomy and serves as a significant visual feature for identifying tumor growth presence or absence. Since the dataset does not have other available metadata such as age, gender, or tumor type, the classification model relies solely on pixel-level features of the images. The features are significant in that they are responsible for capturing minimal intensity variations, textures, and shape deformations found in tumorous tissues.
Before feeding the images into the model, there were several preprocessing steps conducted to enhance data compatibility and quality. First, the images were all resized to a standard resolution of 224 × 224 pixels in order to have uniform input sizes suitable for DenseNet architecture. Then, images were normalized through rescaling pixel values within the range [0, 1], which helps stabilize and accelerate the training process. To prepare the labels for binary classification, one-hot encoding was applied, transforming the categorical labels (‘Tumor’ and ‘No Tumor’) into a machine-readable numeric format.
The dataset was then separated into training and test sets in the proportion of 80:20, and images of 2611 were employed for training purposes and 653 for testing. The split has the benefit of subjecting the model to a vast number of examples while it is being trained as well as evaluating it on unseen data to determine its ability to generalize.
Data pre-processing
Preprocessing is a central step to enhancing model performance, denoising, and ensuring data consistency throughout the pipeline as in Fig. 3. The preprocessing pipeline presented here begins with raw MRI scans, which are normally full of noise, non-brain tissue, and inhomogeneous contrast—meaning preprocessing is unavoidable.

Representation of preprocessing pipeline.
In order to minimize noise, a double filtering approach using Non-Local Means (NLM) and Contrast Limited Adaptive Histogram Equalization (CLAHE) is employed. NLM successfully reduces noise without blurring structural edges, which is extremely crucial for medical images with subtle intensity transitions. CLAHE, on the other hand, enhances local contrast in tumor-susceptible regions without enhancing noise. Through the integration of the two techniques, denoising and contrast enhancement are guaranteed for tumor visibility, resulting in clearer edges surrounding tumor boundaries and hence enhancing feature extraction by DenseNet. After this, morphologically optimized skull-stripping is performed. Traditional skull-stripping procedures run the risk of removing significant brain tissue or even remaining artifacts. In an effort to overcome this, an optimized approach is taken where simple skull-stripping is first performed followed by morphological operations (e.g., closing and opening) to polish the brain mask. This process saves important brain regions and potential tumor waste while eliminating irrelevant non-brain components, maintaining the focus of the model solely on the brain region and not allowing it to learn from irrelevant features. Region-adaptive intensity normalization is applied subsequently. Compared to normal global normalization methods like Z-score or min–max scaling, this approach uses Otsu’s thresholding for segmenting likely tumor and non-tumor areas. Localized histogram normalization is then selectively applied, enhancing contrast in high-probability tumor regions only. This adaptive tumor-aware normalization method enhances the visibility of tumor regions more clearly in deeper convolution layers, which further enhances model performance as well as interpretability, particularly when explainability techniques such as Grad-CAM, IG and LRP are employed.
To improve models’ robustness and generalization, semantic-aware data augmentation is integrated. Instead of standard flips or rotations, this step employs affine-preserving transformations such as shearing and mild distortions that maintain the structural integrity of tumor regions. This ensures the enhanced samples remain clinically relevant without injecting unrealistic patterns when enriching the dataset. Finally, the fully preprocessed image—enhanced, de-noised, and anatomically centered—is passed on to the classification module consisting of DenseNet201 for feature extraction and Support Vector Machine (SVM) for final prediction. This stringently designed pipeline not only improves model performance but also enhances tumor localization, all in the name of improving classification accuracy and model explainability.
The result of preprocessing pipeline on the selected MRI images is a systematic transformation as shown in Figs. 4 and 5 in five phases, each of which is meant to improve visibility of crucial features required in brain tumor diagnosis. Figure 6 shows brain MRI samples demonstrating semantic-aware data augmentation results.

Results of preprocessing pipeline for healthy brain MRI samples.

Results of preprocessing pipeline for brain tumor MRI samples.

Brain MRI samples demonstrating semantic-aware data augmentation results.
Considering Five-Phases technique, the raw grayscale image is the original starting point, typically representing the brain in conjunction with noise, unequal contrast, and non-brain structures like the skull or extra-brain tissues. These raw images tend to be blurry for proper analysis because of inherent noise and contrast limitations.
The second step is Non-Local Means (NLM) denoising, which significantly eliminates granular noise from MRI images. The process smooths out homogenous regions but preserves important anatomical edges such as tumor borders, yielding cleaner images without structural loss. Denoised images appear softer but preserve the structural detail necessary for further processing. During the third stage, local contrast is amplified using Contrast Limited Adaptive Histogram Equalization (CLAHE). CLAHE increases the contrast of low-contrast features by compressing and stretching grayscale levels in the local area without increasing noise. This leads to more uniform brightness and improved visualization of low-intensity regions, useful in enhancing the visibility of tumors that would otherwise have low contrast with the surrounding tissues.
The fourth output, skull stripping, is achieved by using a straightforward thresholding process that masks non-brain regions. Not a clinically correct skull stripping algorithm, this process removes the majority of the skull and background noise, leaving only the brain region. This is significant in that it guarantees that subsequent analysis or models only focus on the region of interest, reducing computational complexity and avoiding false detections outside the brain.
Lastly, the region adaptive enhancement is obtained by using Otsu’s thresholding, which itself automatically identifies a grayscale threshold to separate tumor-like regions from the remainder of the brain. This enhances the most salient regions (which are likely tumors) by destroying smaller regions. The resulting image shows possible pathological regions and prepares data for successful feature extraction or classification, and thus is highly advantageous to utilize for downstream deep learning or radiologic reading.
Collectively, this preprocessing pipeline converts raw MRI inputs into progressively more accurate representations, rendering them progressively more susceptible to automated identification of brain tumors and facilitating clinical diagnosis.
Model selection
Once the data is ready, the second most critical step is to select and apply the hybrid classification architecture. The model applied in this study is a combination of DenseNet—a densely connected convolutional neural network—and a Support Vector Machine (SVM), combined further with the Grad-CAM explainability system39. Each component in this architecture has a unique and complementary function in the system as a whole.
The DenseNet architecture, proposed by Gao Huang et al.40, is employed for the extraction of deep features from the MRI scans. DenseNet possesses one of the main strengths that its pattern of connectivity is such that each layer receives input from all previous layers. Dense, connected architecture facilitates effective reuse of features, improves the gradient flow through backpropagation, and enables the training of deeper networks without vanishing gradients issues. Thus, DenseNet is able to learn intricate spatial information and texture patterns required for distinguishing tumor regions from normal tissue in brain imaging.
The feature vectors extracted by using DenseNet are then passed to a linear Support Vector Machine classifier41. SVM performs well in high-dimensional space and is also able to find the optimal hyperplane that distinguishes various classes with maximum margin. In the instance of this study, the SVM classifier determines whether a given MRI image is tumor or non-tumor using the learned features. The hybrid approach enhances classification performance using DenseNet’s robust feature extraction and SVM’s robust decision boundary formation.
To increase the explainability of the model’s decisions and make the predictions interpretable to medical practitioners, the model employs Gradient-weighted Class Activation Mapping (Grad-CAM)42. Grad-CAM is an XAI method that generates heatmaps to represent those regions of the image that had the greatest impact on the final prediction. By overlaying these heatmaps on the real MRI scans, physicians can gain a greater understanding of what regions of anatomy the model focused on when making its determination. This explanation visualization serves to bridge the gap between machine-based predictions and acceptance in the clinic, making the system more applicable for implementation into real healthcare settings.
Model evaluation
Performance evaluation of the proposed model is a critical step to determine if the model can be employed for brain tumor diagnosis or not. Statistical metrics are utilized to evaluate the model’s classification accuracy on the test set. The foundation of these metrics43 is the confusion matrix, which presents a summary of the model’s true positive, true negative, false positive, and false negative predictions.
From Table 1’s confusion matrix, some of the indicators of evaluation are derived. Accuracy, calculated as the proportion of correct predictions to the total number of instances, gives the overall picture of model performance. Sensitivity or recall attempts the ability of the model to actually correctly classify tumor cases, which matters much in medicine where false negatives are extremely expensive. Specificity refers to the accuracy with which the model is able to identify healthy cases, so as not to raise false alarms. Precision assesses the proportion of actual tumor cases out of all the cases the model predicts as positive, and this is a measure of precision of the model. Finally, the F1-score, the harmonic mean of precision and recall, provides a balanced score that considers both false positives and false negatives.
Together, these steps44 form a comprehensive assessment framework to ensure that the model not only performs well in terms of predictive accuracy but also clinical reliability and validity up to high levels. The ultimate aim is to ensure that the system developed is not only technically valid but also clinically relevant, improving improved diagnostic protocols and patient care. We applied a hybrid deep learning approach for brain tumor classification from MRI scans of the Kaggle dataset acquired by Sartaj in this study. The dataset consists of an evenly distributed set of labeled brain MRI scans as tumor and normal brain MRI types to present a good estimation of classification performance. Images were also resized to 224 × 224 pixels and normalized as per DenseNet201’s input requirements.
Feature extraction used DenseNet201, a deep learning model pre-trained on dense connectivity that helps facilitate the flow of gradients and alleviates vanishing gradient risk. The model was initialized with ImageNet weights and further fine-tuned using the Kaggle dataset. Deep features were obtained from the global average pooling layer of DenseNet201, which identified high- dimensional representations of tumor-associated patterns. The features extracted were then fed into a Support Vector Machine (SVM) classifier with an RBF kernel, which performed better in classifying tumor and non-tumor cases. Hyperparameters were optimized, with regularization strength C = 1.0 and gamma = 0.01 to achieve a balance between model complexity and generalizability.
To enable explainability, Grad-CAM (Gradient-weighted Class Activation Mapping) was incorporated as the primary XAI technique. Grad-CAM was used to visualize the most influential regions in MRI scans that contributed to the model’s predictions, thus enhancing interpretability for physicians. The last convolutional layer of DenseNet201 was used for gradient-based activation mapping, producing heatmaps highlighting the tumor-infected regions in MRI scans. This can help clinicians with an open understanding of model decisions, overcoming the black-box nature of deep learning models in medical diagnostics.
It was trained for 50 epochs using the Adam optimizer with a learning rate of 0.0001 and batch size of 32. Training was performed on an NVIDIA RTX 3090 GPU, which possesses 24 GB VRAM, significantly reducing computation time and allowing high-resolution MRI images to be processed efficiently. Evaluation was done based on conventional evaluation metrics, including accuracy, precision, recall, F1-score, and specificity, to effectively measure model performance. The combination of DenseNet201 for feature extraction, SVM for classification, and Grad-CAM for explainability formed a high-performance and interpretable AI-driven diagnostic tool for brain tumor classification.
link